How to Select Antibodies and Phosphorylated Antibodies for WB?
In Western Blot (WB) experiments, selecting appropriate antibodies is key to ensuring experimental success.
1. Core Selection Principle: Validation First
Before purchasing antibodies, absolutely do not only look at the catalog number or price; the first thing to look at is the antibody validation data.
The first pitfall that beginners are most likely to step into: blindly selecting antibodies labeled for immunoassays, ignoring whether they have been validated for WB experiments. Different experimental methods have different requirements for antibody epitope recognition. For example, antibodies suitable for IHC may recognize the native conformation of proteins, while in WB experiments, samples undergo SDS denaturation and boiling, destroying protein conformation, and only linear epitopes can be recognized—if the antibody has not been validated for WB, even if the target matches, there may be no signal or excessive non-specific bands.
2. Confirm Target Sequence
Step 1: Clarify core information of the target protein: First, use databases such as Uniprot and NCBI to query the official gene symbol, amino acid sequence, molecular weight, structural domains, and other information of the target protein, and confirm the species source and subtype of the target protein.
Step 2: Analyze specific regions of the target protein: Antibodies recognize protein "epitopes" (antigenic determinants), which are divided into linear epitopes and conformational epitopes. In WB experiments, after sample denaturation, only linear epitopes can be recognized, so antibodies targeting linear epitopes need to be selected. At the same time, avoid conserved sequence regions of the target protein and homologous proteins, and select specific epitopes. For example, if the target protein is a member of a protein family, you can compare the amino acid sequences of each member of the family through databases, find the amino acid fragment unique to the target protein, and select antibodies targeting that fragment to reduce non-specific bands from the root.
Step 3: Adjust sequence selection based on experimental purpose: If the experimental purpose is to detect the total expression of the target protein, you can choose antibodies targeting the constant region of the protein; if the experimental purpose is to detect splice variants or mutant forms of the protein, you need to select antibodies targeting the regions containing splice sites or mutation sites to ensure only the target form of the protein is recognized.
3. Sequence and Site Selection for Phosphorylation Antibodies
Selection logic for phosphorylation sites: Protein phosphorylation sites are usually located on serine (Ser), threonine (Thr), and tyrosine (Tyr) residues. Phosphorylation at different sites corresponds to different signaling pathway activation states. Therefore, before selecting phosphorylation antibodies, you need to first clarify the experimental purpose: which signaling pathway activation is being investigated, and then determine the key phosphorylation sites of the target protein. For example, the core activation sites of AKT protein are Ser473 and Thr308. If you are studying apoptosis, cell cycle, cytoskeletal reorganization, or cancer-related topics, both Thr308 and Ser473 sites may be involved; if studying cell differentiation, you may need to pay more attention to the Ser473 site; protein degradation-related studies generally involve the Ser473 site; in non-small cell lung cancer research, it may be highly related to phosphorylation at the Thr308 site.
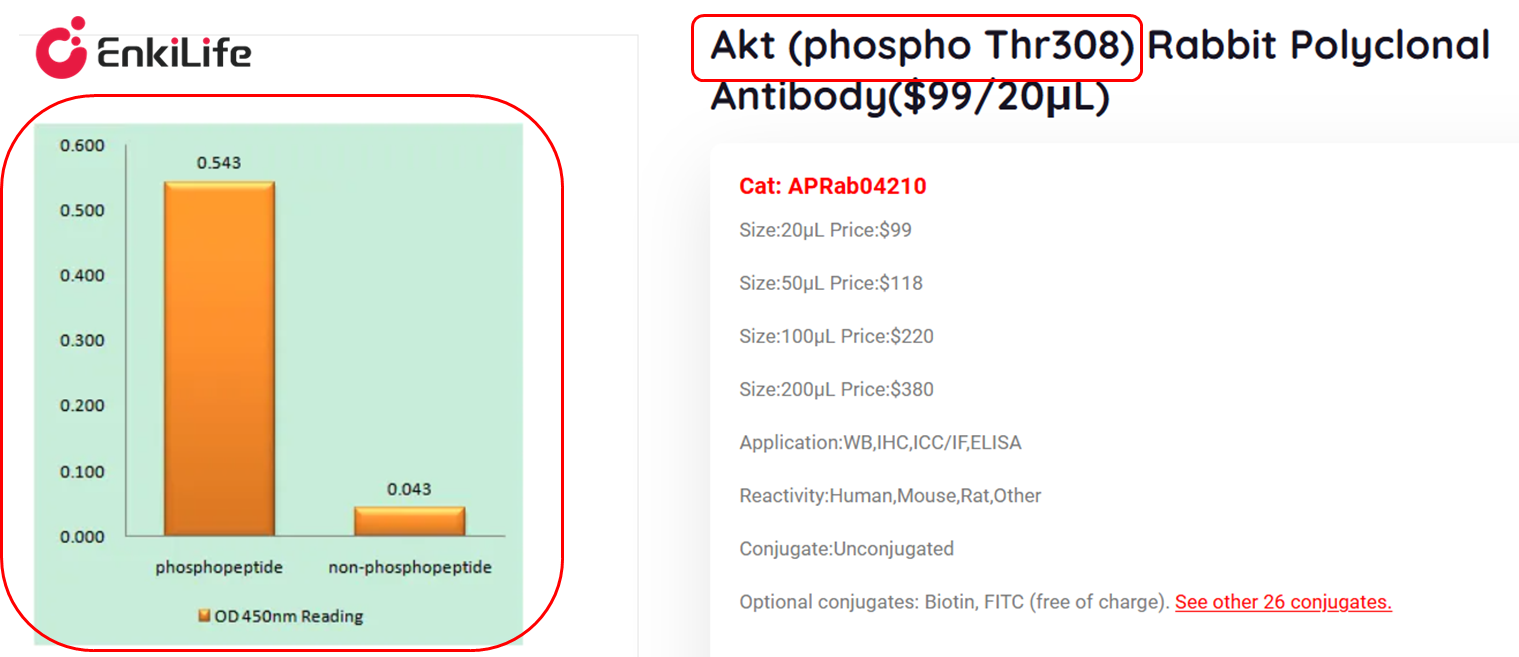
How to confirm antibody site specificity: Check the antibody specification, which must clearly label the phosphorylation site and indicate that it recognizes only the phosphorylated form. If the specification does not clearly label the site, or labels recognition of phosphorylated Ser/Thr/Tyr residues without specific site information, then this antibody is a "pan-phosphorylation antibody," which is only suitable for screening unknown phosphorylation sites and not suitable for specific detection of phosphorylation status at a particular site. Beginners should avoid using this type of antibody for precise detection.
Sequence adaptation considerations for phosphorylation antibodies: The epitope of phosphorylation antibodies not only contains the phosphorylation site but also the amino acid sequence around the site. This part of the sequence determines the antibody specificity—for example, two antibodies targeting p-AKT Ser473, if their recognized epitopes contain different amino acid sequences around Ser473, may cause one of the antibodies to cross-react with phosphorylation sites of other homologous proteins, resulting in non-specific bands. Therefore, when selecting, you need to check the antigen sequence in the specification to confirm that it contains the target phosphorylation site and the surrounding specific sequence. At the same time, you can check the literature to confirm whether this antibody has cross-reactivity in similar experiments.
Avoid common misconceptions about phosphorylation antibodies: ① Confusing phosphorylation antibodies with total protein antibodies: Total protein antibodies recognize all forms of the target protein, including phosphorylated and unphosphorylated, while phosphorylation antibodies only recognize the phosphorylated form at specific sites. The two cannot substitute for each other, and both types of antibodies are usually needed in experiments, using total protein expression as an internal reference to eliminate interference from uneven loading; ② Ignoring the conservation of phosphorylation sites: If experimental samples are cross-species, such as using human-derived antibodies on mouse samples, you need to confirm that the target phosphorylation site is conserved in that species, otherwise the antibody cannot recognize it; ③ Blindly selecting pan-phosphorylation antibodies: If beginners have not identified the target phosphorylation site, do not arbitrarily use pan-phosphorylation antibodies. This type of antibody has poor specificity and is prone to false positives. It is recommended to first determine key phosphorylation sites through literature or preliminary experiments, then select specific antibodies for the corresponding sites.
4. Species Reactivity Matching
Strictly match sample species: Confirm the Reactivity (species reactivity) column in the antibody specification, clearly labeling the species of the experimental sample. For example, human samples require antibodies labeled as Human, and mouse samples require antibodies labeled as Mouse. Do not arbitrarily use across species—for example, rabbit anti-human p-AKT Ser473 antibody may not recognize p-AKT Ser473 in mouse samples. Even if the sequence homology is high, recognition failure may occur due to amino acid differences.
Considerations for cross-species use: If you truly need to use antibodies across species in experiments and there are no specific antibodies for the corresponding species, two conditions must be met: ① The specification clearly labels "Cross-reactivity," for example labeling Reactivity: Human, Mouse, indicating that the antibody can simultaneously recognize the target proteins of both human and mouse; ② Check the literature to confirm that colleagues have successfully used this antibody across species, and verify through preliminary experiments, such as setting up positive controls to confirm the presence of bands without obvious non-specific bands. For phosphorylation antibodies, cross-species use requires additional confirmation that the target phosphorylation site is conserved in both species, otherwise the antibody will not be able to recognize it.
5. Antibody Source and Form Adaptation
Common sources: In WB experiments, rabbit-derived and mouse-derived antibodies are most commonly used, each with their advantages. Rabbit-derived antibodies have high affinity and strong specificity, suitable for detecting low-abundance proteins; mouse-derived antibodies have good stability and small batch-to-batch variation, suitable for quantitative WB experiments. Beginners can prioritize rabbit-derived antibodies, which have broader applicability; if experiments require high quantitative accuracy, mouse-derived monoclonal antibodies can be selected.
Monoclonal antibodies: Produced by a single B-cell clone, they recognize only one linear epitope of the target protein, with extremely high specificity, few non-specific bands, and small batch-to-batch variation. They are suitable for quantitative WB and phosphorylated protein detection—because phosphorylation antibodies need to precisely recognize specific sites, monoclonal antibodies can effectively avoid cross-reactivity with other sites and homologous proteins, making them the preferred form for phosphorylation antibodies.
Polyclonal antibodies: Produced by multiple B-cell clones, they can recognize multiple epitopes of the target protein, with high sensitivity and strong binding ability, suitable for detecting low-abundance total proteins, but with relatively lower specificity and prone to non-specific bands—beginners are not recommended to choose polyclonal antibodies for phosphorylated protein detection unless there are no corresponding monoclonal antibodies, and conditions need to be optimized through preliminary experiments to reduce non-specific bands.
Key reminder: After WB samples undergo SDS denaturation, protein conformation is destroyed and only linear epitopes can be recognized. Therefore, regardless of which antibody you choose, you must avoid selecting antibodies that only recognize native conformation—this type of antibody cannot recognize denatured proteins and will result in no signal. Phosphorylation antibodies especially need to pay attention to this point, because phosphorylation epitopes themselves are relatively fragile, and if the antibody recognizes a conformational epitope after denaturation, it will directly lose recognition ability.
